MCMC sampling for Bayesian quantile regression with an unknown (nonparametric) transformation. Like in traditional Bayesian quantile regression, an asymmetric Laplace distribution is assumed for the errors, so the regression models targets the specified quantile. However, these models are often woefully inadequate for describing observed data. We introduce a nonparametric transformation to improve model adequacy while still providing inference for the regression coefficients and the specified quantile. A g-prior is assumed for the regression coefficients.
Usage
sbqr(
y,
X,
tau = 0.5,
X_test = X,
psi = length(y),
laplace_approx = TRUE,
fixedX = TRUE,
approx_g = FALSE,
nsave = 1000,
nburn = 100,
ngrid = 100,
verbose = TRUE
)Arguments
- y
n x 1response vector- X
n x pmatrix of predictors (no intercept)- tau
the target quantile (between zero and one)
- X_test
n_test x pmatrix of predictors for test data; default is the observed covariatesX- psi
prior variance (g-prior)
- laplace_approx
logical; if TRUE, use a normal approximation to the posterior in the definition of the transformation; otherwise the prior is used
- fixedX
logical; if TRUE, treat the design as fixed (non-random) when sampling the transformation; otherwise treat covariates as random with an unknown distribution
- approx_g
logical; if TRUE, apply large-sample approximation for the transformation
- nsave
number of MCMC iterations to save
- nburn
number of MCMC iterations to discard
- ngrid
number of grid points for inverse approximations
- verbose
logical; if TRUE, print time remaining
Value
a list with the following elements:
coefficientsthe posterior mean of the regression coefficientsfitted.valuesthe estimatedtauth quantile at test pointsX_testpost_theta:nsave x psamples from the posterior distribution of the regression coefficientspost_ypred:nsave x n_testsamples from the posterior predictive distribution at test pointsX_testpost_qtau:nsave x n_testsamples of thetauth conditional quantile at test pointsX_testpost_g:nsaveposterior samples of the transformation evaluated at the uniqueyvaluesmodel: the model fit (here,sbqr)
as well as the arguments passed in.
Details
This function provides fully Bayesian inference for a
transformed quantile linear model.
The transformation is modeled as unknown and learned jointly
with the regression coefficients (unless approx_g = TRUE, which then uses
a point approximation). This model applies for real-valued data, positive data, and
compactly-supported data (the support is automatically deduced from the observed y values).
The results are typically unchanged whether laplace_approx is TRUE/FALSE;
setting it to TRUE may reduce sensitivity to the prior, while setting it to FALSE
may speed up computations for very large datasets. Similarly, treating the
covariates as fixed (fixedX = TRUE) can substantially improve
computing efficiency, so we make this the default.
Note
The location (intercept) is not identified, so any intercepts
in X and X_test will be removed. The model-fitting *does*
include an internal location-scale adjustment, but the function only outputs
inferential summaries for the identifiable parameters.
Examples
# \donttest{
# Simulate some heteroskedastic data (no transformation):
dat = simulate_tlm(n = 200, p = 10, g_type = 'box-cox', heterosked = TRUE, lambda = 1)
y = dat$y; X = dat$X # training data
y_test = dat$y_test; X_test = dat$X_test # testing data
# Target this quantile:
tau = 0.05
# Fit the semiparametric Bayesian quantile regression model:
fit = sbqr(y = y, X = X, tau = tau, X_test = X_test)
#> [1] "4 sec remaining"
#> [1] "2 sec remaining"
#> [1] "0 sec remaining"
#> [1] "Total time: 6 seconds"
names(fit) # what is returned
#> [1] "coefficients" "fitted.values" "post_theta" "post_ypred"
#> [5] "post_qtau" "post_g" "model" "y"
#> [9] "X" "X_test" "psi" "laplace_approx"
#> [13] "fixedX" "approx_g" "tau"
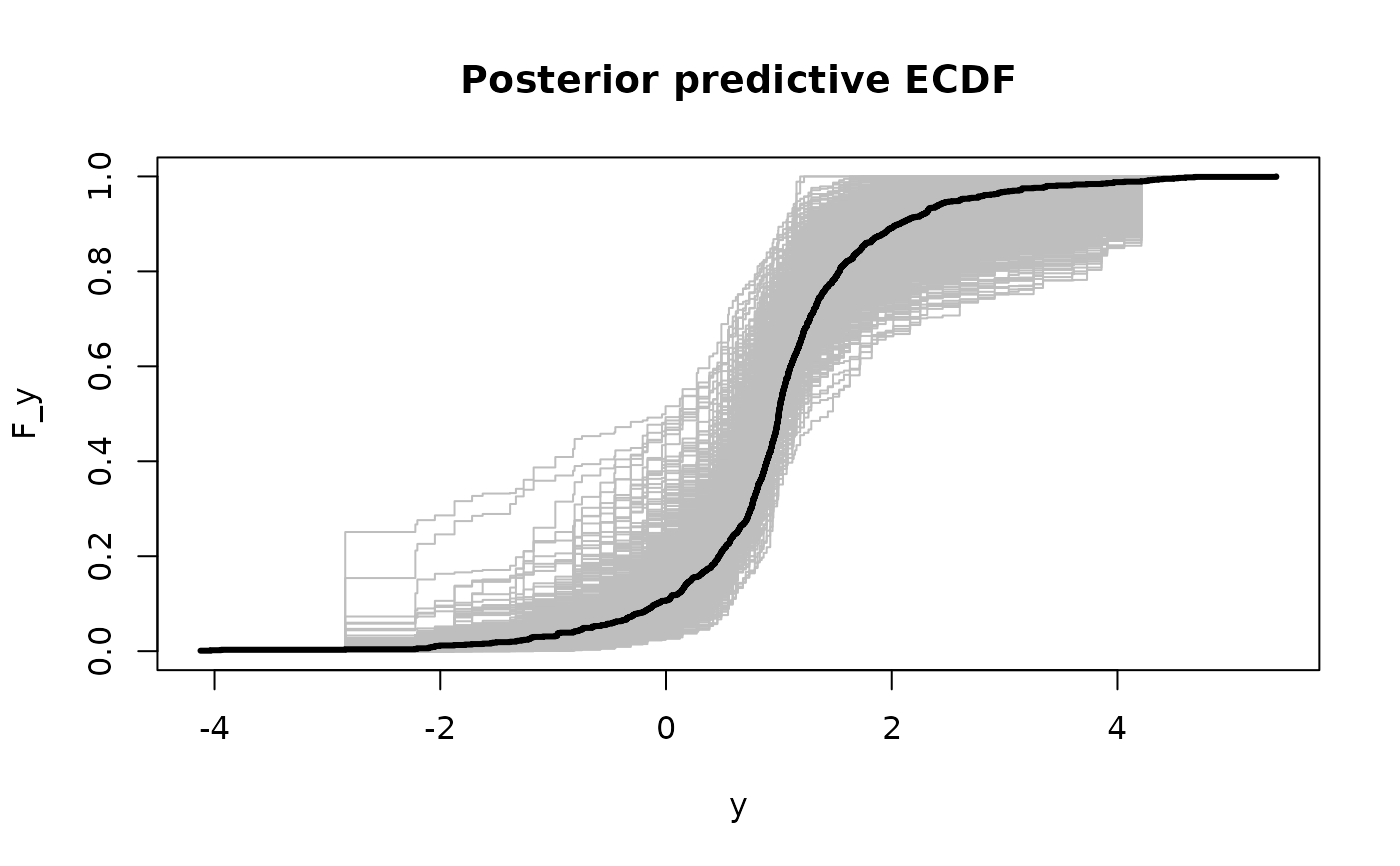
# Posterior predictive checks on testing data: empirical CDF
y0 = sort(unique(y_test))
plot(y0, y0, type='n', ylim = c(0,1),
xlab='y', ylab='F_y', main = 'Posterior predictive ECDF')
temp = sapply(1:nrow(fit$post_ypred), function(s)
lines(y0, ecdf(fit$post_ypred[s,])(y0), # ECDF of posterior predictive draws
col='gray', type ='s'))
lines(y0, ecdf(y_test)(y0), # ECDF of testing data
col='black', type = 's', lwd = 3)
 # }
# }